Dr. Boris Zakharov
For the last 25 years, I’ve studied ground spiders (Araneae, Gnaphosidae) as a member of the Arachnological group of the Invertebrate Zoology Department at American Museum of Natural History in New York. My primary interest lies in studying spider taxonomy. Taxonomy is a scientific discipline preoccupied with the discovery of new species and classification of organisms. It is probably the oldest and most ancient scientific discipline. Why did I choose it? Well, I did so because taxonomy and its theoretical foundation, classification, are fundamental tenets of Biology (and, actually, not only Biology, but a great deal of scientific knowledge in general). To classify species, a scientist has to answer the following fundamental questions: (1) What are species?, (2) What basic principles (ideas or theories) can we utilize so as to fundamentally classify organisms?, (3) What actually allows for the classification of organisms? To answer these questions, taxonomists have to review a broad scope of knowledge in organisms’ morphologies (structures), molecular biologies, ecologies, and evolutions. Altogether, they make the theory of classification the most fundamental and general discipline that can provide a comprehensive organizational scheme to all of contemporary biological knowledge.
The American Museum of Natural History has the largest collection of ground spiders from all over the world. Spiders belong to the largest order in the class of Arachnida. At present time, the order includes 48,440 species and 120 families, and the list of spiders is still far from being complete. Ground spiders (family Gnaphosidae) are among the biggest and most diverse groups of spiders. They are classified across 158 genera and 2,522 species, and their taxonomy is still in the process of active development. New achievements in DNA analysis, the ability to process vast amounts of data, and new imaging techniques have dramatically changed taxonomic research in the last few decades. Now, a taxonomist has to operate not only with morphological data, but master DNA analysis and tinker with computer programs for evolutionary reconstructions. Incredible advances have occurred in the implementation of imaging techniques in recent years. Just 50 years ago, the most common presentation of morphologic organization of living organisms was a scientific drawing, and so a taxonomist then was not only specialist in understanding a particular group of animals, but was also an expert artist! Now, most illustrations that demonstrate specific characteristics of species are made with digital cameras, electron microscopes, and/or CT Scanners.

Advanced imaging techniques have paved the way for new opportunities in the study of Anatomy and Functional Morphology. Usually, when we think about CT or MRI scans, what comes to our mind is a laboratory room somewhere in a hospital, where medical specialists examine patients’ brains or organs residing in the abdominal cavity. However, applications of these techniques have come quite far in the information they can reveal. CT scanners are widely used in various disciplines for the study of fossils, non-invasive study of microscopic anatomy of invertebrates, functional anatomy of muscle contraction, etc.
I began my micro-computed tomography studies in 2015 in the Image Laboratory at the American Museum of Natural History (MIF) together with my former science student, Andrius Jankauskas. At that time, he was actually a student at the Engineering Department of City College of CUNY. We used X-ray micro-computed tomography to visualize minute anatomic organization in ground spiders.

CT scanning consists of the construction of images by exposing an object of interest to X-rays. This technique works perfectly for solid, hard objects such as bones, fossils consisting of materials with different density, etc. Spiders, whose organs are made of soft tissues, pose a challenge in that regard. Their soft tissues are highly transparent for X-rays and, in light of this, images created via the use of X-rays are very blurred and exhibit difficult-to-find borders. Also, increasin the intensity of X-ray radiation does not help much to improve image resolution nor image contrast. Histologists, scientists who study tissues, learn how to make different soft tissues visible. In the process, they stain such tissues with different colors. We also stain our spiders, and this is achieved by employing different agents such as iodine to obtain high-contrast images of soft tissues and muscles. Spiders with their soft bodies seem to be ideal objects to undergo this kind of treatment.
The preparation of a spider specimen for scanning takes a few days. Spiders in the AMNH collection are preserved in 75% alcohol. A few days before imaging, we clean our specimen in fresh alcohol by ultrasound. Vibrations of fluids generated by ultrasound remove small contaminant particles from a spider’s body. After the use of ultrasound, we hold the specimen in 95% alcohol for 24 hours. 95% alcohol removes water molecules from the soft tissues of a spider’s body. We are then ready to stain our spider of interest. We place the spider in stain, and hold it there for some time depending on what kind of stain we use. For example, in the case of iodine, we leave the specimen overnight. Different stains have different degrees of affinity for different tissues, and so you can manipulate them to contrast the tissue of our interest: muscle, nerve, or connective tissue. Our stained specimen is then mounted on a prepared stage that does not allow for drying and moving. Now, we are ready to scan our spider!
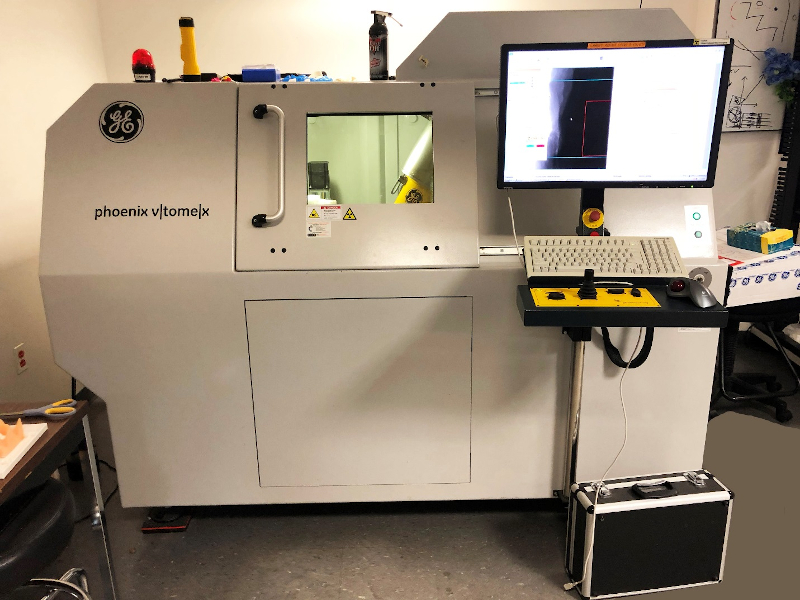
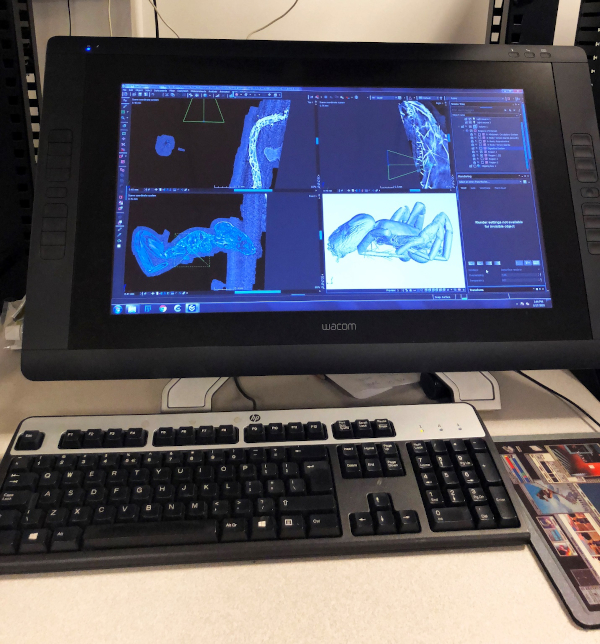
We carried out our X-ray scans on the GE VTOMEX 240 X-ray 3D CT Scanner (see Fig. 2) in the Microscopy and Imaging Facility (MIF) of the American Museum of Natural History. Our prepared specimen was inserted into a chamber with a X-ray emitting scanner. The process of scanning involved taking a series of images. In order to achieve this, the stage upon which our specimen was mounted rotated about the vertical direction (Z direction). The scanner, in turn, made X-ray images of the specimen at different angles. These images appeared as spots of varying intensity using gray scale and with any one spot signifying the density of a specific point on the specimen. The slides used were organized in 3 stacks along the 3 directions (X, Y, and Z) of 3D space and then processed with the software ‘VGStudio Max 3.0,’ which accepted the generated image stacks as raw input. Instead of working with vector values like in most 3D CAD software, this particular program uses voxel (pixel in 3D space) geometry. The program allows one to construct a region of interest in 2D space of the image stack representation, with all three axis interpolated, and to display 3D objects with their specified transparencies and density ranges. Such capabilities enabled us to inspect the spider specimen under invesitgation and rebuild its internal structures, which can further be exported as images (see Fig. 3), video, or 3D vector graphics (for animation purposes or 3D printing).
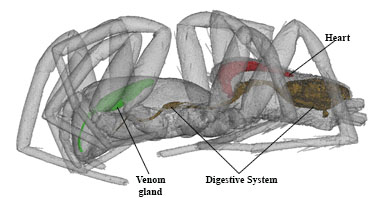
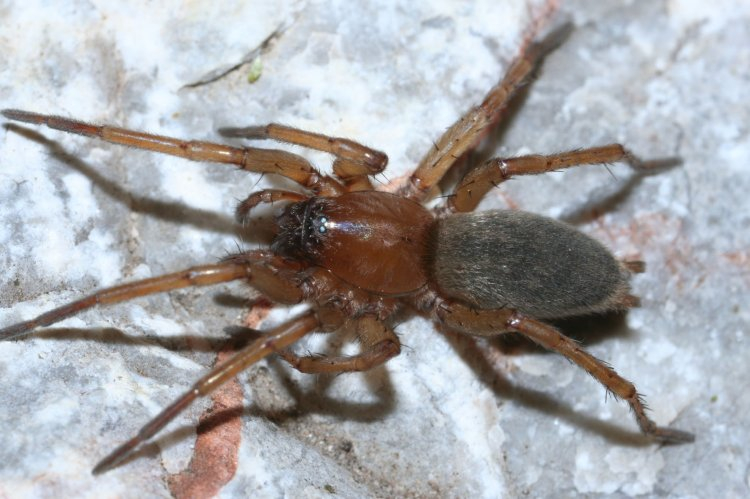
Stained tissues form features of different density. This helps us to pinpoint specific organs, their internal structures, and systems. Reconstructed images are ultimately processed, combined, represented in color, and labeled with the Photoshop CS6 computer program as shown below in Fig. 3.